
Welcome to MZmol
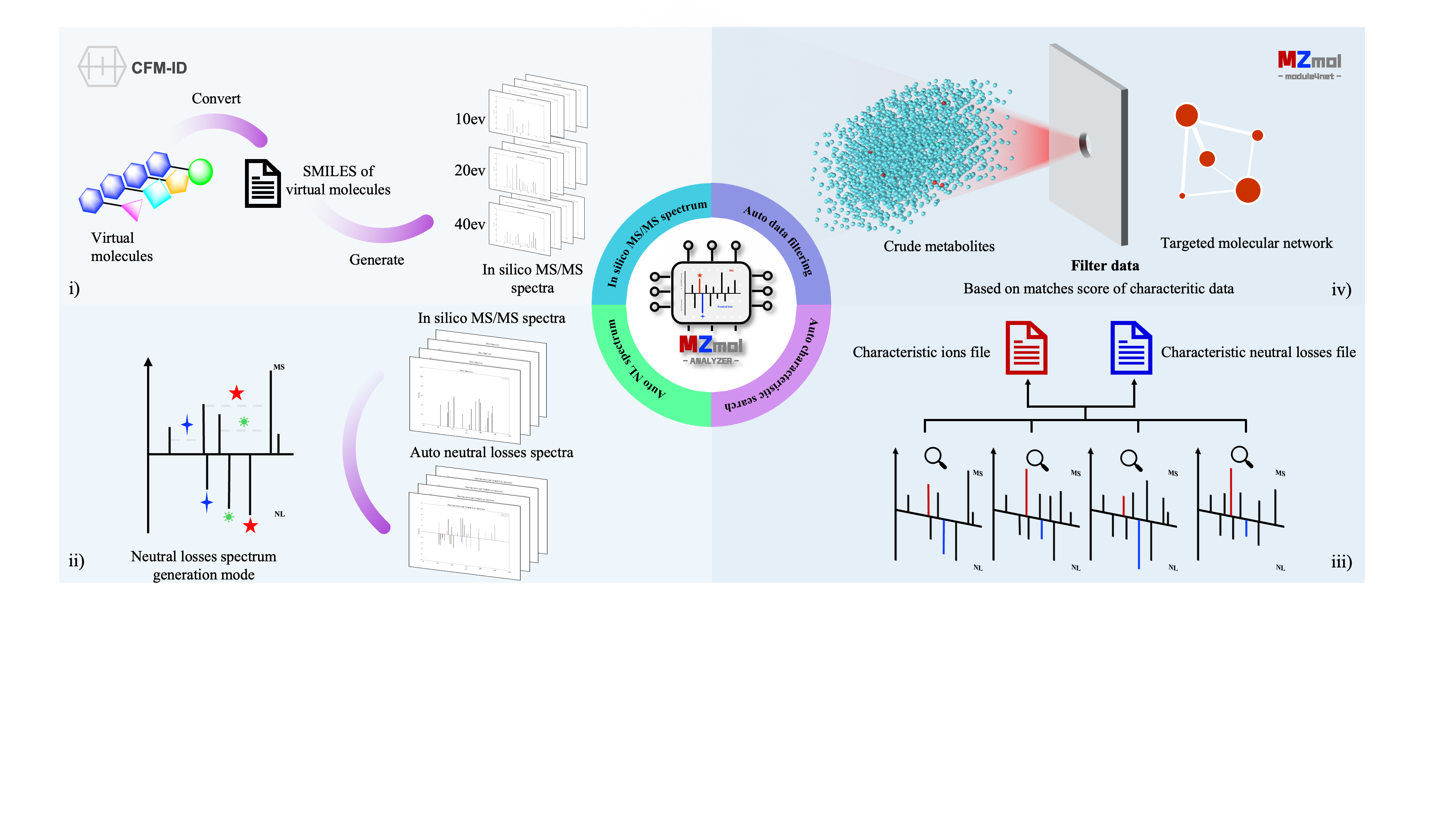
MZmol is an open-source online service platform designed to assist researchers in easily utilizing in-silico MS/MS data. It automates the generation of:
- Neutral loss mass spectra
- Characteristic mass spectrometry data
- Filtering of targeted metabolite networks
This enables the identification and isolation of unknown compounds from unknown metabolites, a strategy we refer to as the Virtual Molecular Network (SMMN) approach.
Current Technology
Currently, MZmol utilizes the open-source CFM-ID 4.0 model for generating in-silico MS/MS data. As the precision of simulated mass spectrometry data continues to improve, more advanced models will be incorporated in future updates, further enhancing compound mass spectra prediction capabilities.
Citation
When using MZmol, please cite the following reference:
Peng Y, Luo S, Li Q, Li Y, Zhang J, Ding N, Sun W, Chen C, Liu J, Ye Y, Zhu H, and Zhang Y. (2025).
Discovery of Indole Alkaloids Crienamides A and B from Penicillium citrinum by a Simulated MS/MS-Guided Molecular Network Strategy.
Org Lett.ASAP
Wang F, Liigand J, Tian S, Arndt D, Greiner R, and Wishart D. (2021).
CFM-ID 4.0: More Accurate ESI MS/MS Spectral Prediction and Compound Identification.
Anal Chem. 93(34):11692-11700.
As the platform evolves, it will provide even more advanced tools to help researchers utilize MZmol more effectively in their studies.